Hello,
I’m fairly new to Psi4 and I’m trying to calculate ESP values on a grid surrounding a molecule. I’m comparing in-memory methods and out of memory methods, and trying to figure out why I’m seeing different values.
Setup:
import psi4
import numpy as np
import psi4.core as p4c
mol_str = """ C 3.9336793405012425 0.5346774188749415 -0.8928514517978514
C 5.303094945924056 0.5873177581665312 -1.159691036377977
C 6.201504049592222 -0.09149376798441175 -0.3402856747330883
C 5.733931689556879 -0.8248754888890404 0.7471407426068619
C 4.365772298366927 -0.8805636264491232 1.0179806816982493
C 3.445952574452895 -0.20476876458844664 0.1991302049836341
C 1.9654092347012313 -0.2544474458229271 0.5310879493895444
C 1.1029097243138335 -0.6323124209287291 -0.6838098911484615
C -0.1923054043763034 -1.2195732040662057 -0.17913424960688004
O -0.19156427004666907 -2.18144179203173 0.5818630559291688
N -1.3234892209028375 -0.5346914074513771 -0.5785423504115056
C -2.392512269533669 -0.3441914256151788 0.45023232707137256
C -3.757087918366187 0.01515122013256527 -0.21053784730799055
C -4.887067134630283 0.02948733841113557 0.8392867273640869
C -4.1232090625744275 -1.0532693834407665 -1.2646772156805537
C -3.7167896842382344 1.3915644978122723 -0.9006784879882505
C -1.892420255349704 0.7024667920178694 1.521741420529058
O -0.8312916013131516 1.3340723435394133 1.1669477652581894
O -2.4911362558218046 0.7657895117293508 2.619048850338715
N 1.5600250520198258 1.0569732117075195 1.088566166274743
H 3.26044332755187 1.0744074733250508 -1.5557806902345677
H 5.672724253339586 1.155753762822649 -2.0105624319650426
H 7.267808336298617 -0.05245107036434195 -0.5513108674032534
H 6.4372820073529535 -1.3577900731669172 1.3836055722598994
H 4.0295174232231155 -1.46604718945525 1.8731631007255978
H 1.8090111290385975 -0.9667139389635266 1.3495857786410308
H 1.5842900491058522 -1.4131018866310743 -1.2858510769143463
H 0.9311687283330116 0.22616804138985505 -1.3424029434864222
H -1.148244857222129 0.33853262127457845 -1.0573389738533145
H -2.4965031136449802 -1.2997662388946127 0.9822052241022017
H -4.904558921288868 -0.9025206479949446 1.4149829124728843
H -4.783612836357871 0.8627561316859041 1.540902950328714
H -5.8678293602966445 0.1407769552406649 0.3618274510955099
H -4.143572308782679 -2.0543600311298205 -0.8193547135461033
H -5.112731058086654 -0.8615793144608911 -1.695781358871089
H -3.409022406295142 -1.0679358709617324 -2.0950322631427007
H -3.479455283242055 2.191090518556378 -0.19094454680130502
H -4.687447951040984 1.632507493639407 -1.3498526235501647
H -2.9729773947805813 1.4221109747734484 -1.7034978805320657
H 0.5093459150911678 1.1241882218928128 1.2018998320578946
H 1.7838446160867416 1.864607897901198 0.5082946616512665
H 1.907113873342181 1.2134948043971896 2.0364772615019517"""
mol = psi4.geometry(mol_str)
mol.update_geometry()
x = np.linspace(-7, 8, 15)
y = np.linspace(-3, 3, 15)
z = np.linspace(-3, 2.5, 15)
out = np.meshgrid(x,y,z)
points = np.stack(out).reshape(3,-1).T
Out of memory:
np.savetxt('grid.dat', points, fmt='%15.10f')
psi4.core.set_active_molecule(mol)
psi4.set_options({'basis': '6-31g*'})
psi4.set_options({})
psi4.prop('scf', properties=['GRID_ESP'])
psi4.core.clean()
grd_om = np.loadtxt('grid_esp.dat').reshape(15,15,15)
In memory:
ene, wfn = psi4.energy('scf/6-31g*', return_wfn = True)
myepc = p4c.ESPPropCalc(wfn)
psi4_matrix = p4c.Matrix.from_array(points)
esps = np.array(myepc.compute_esp_over_grid_in_memory(psi4_matrix))
grd_im = esps.reshape(15,15,15)
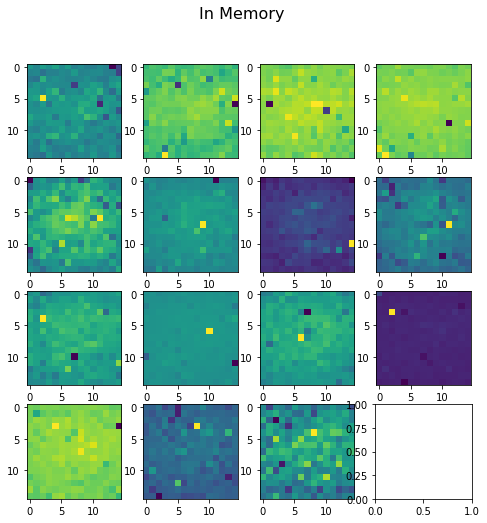
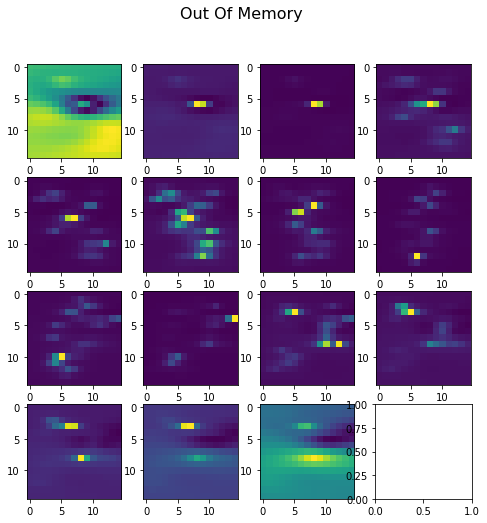
Both methods generate the same energy value of -914, but the values of the potentials calculated on the grid points is drastically different. For the in memory grid computation, values range from -194 to 480. For the out of memory grid computation, values range from -0.14 to 20.
Here’s visualizing the difference plotting slices of the solved grid:
Is there some setting I’m missing?